Date: Tue, 25 Aug 2020 15:24:29 +0300
Hello!
I'm trying to investigate a polyglutamilation effect in tubulins (
https://en.wikipedia.org/wiki/Polyglutamylation or
https://www.nature.com/articles/s41594-020-0462-0#Sec25 ), which seems to
be a slightly tricky task.
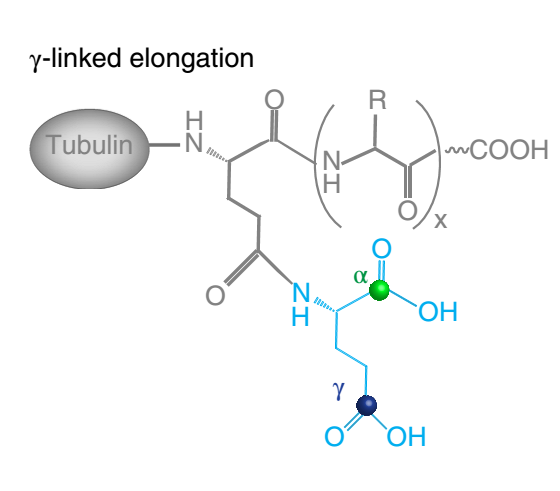
First of all I've to generate a non-terminal glutamine which is a part of
the chain (for example, chain A), but its terminal gamma carboxyl group
forms a peptide bond with glutmaine tag (in attachment). I think, that I
can build such strange amino acid, forming three peptide bonds at once,
with R.E.D.III or something else. Thus, it should look like a simple
in-chain glutamine with a replaced gamma-carboxyl oxygen by amide group of
the connected glutamine. This is already difficult manipulation, but the
number of the added glutamines can vary.
Another way to generate a new topology based on GAFF force field
parameters library. As we can replace carboxyl group with amide group of
asparagine and calculate the difference in charges with gaussian/gamess.
Is it real or not?
Then I thought about the possibility of binding together of two separate
chains (like cys-cys) - namely, one chain contains a modified glutamine
residue, and all substituted in gamma position glutamines belong to another
chain. Is it possible or not? Of course charge distribution should be
correctly tuned up to reflect natural behaviour of the construction.
Is it possible or not? Do You have any suggestions?
Thank You!
[image: glu.png]
*Nemo me impune lacessit*
[image: Mailtrack]
<https://mailtrack.io?utm_source=gmail&utm_medium=signature&utm_campaign=signaturevirality5&>
Sender
notified by
Mailtrack
<https://mailtrack.io?utm_source=gmail&utm_medium=signature&utm_campaign=signaturevirality5&>
25.08.20,
15:05:58
_______________________________________________
AMBER mailing list
AMBER.ambermd.org
http://lists.ambermd.org/mailman/listinfo/amber
(image/png attachment: glu.png)
