Date: Mon, 10 Jan 2022 13:59:19 -0500
Hi Amber user
I am working on a ligand that is covalently bound to FAD cofactor. . *Both
PO4s meioties of FAD together have -2 whereas the drug part has -1 (please
see par.pdb). *Overall charge on this FAD bound drug is -1.
To prepare this compound, i used this command
*antechamber -i par.pdb -fi pdb -o lig.am1bcc.mol2 -fo mol2 -c bcc -s 2
-nc -1*
*but the output lig.am1bcc.mol2 shows 0 charge and two PO4 meioties have
also changed. *
*In par.pdb ligand structure, P atom has two oxygen (O1P and O1P), O1P has
-1 charge and valency of O2P is complete and shows double bonds. Same is
the case with PA oxygens. In lig.am1bcc.mol charges on thes PO4 atoms and
drug have been removed and overall charge is zero (please see
lig.am1bcc.mol2 file).*
- *In another attempt I tried parameterizing it with sybyl and gaff2.
Output files were even worse. *
* antechamber -i par.mol2 -fi mol2 -o ligand-out.mol2 -fo mol2
-at sybyl -pf y*
* antechamber -i ligand-out.mol2 -fi mol2 -o lig.am1bcc.mol2
-fo mol2 -at gaff2 -c bcc -rn LIG -pf yes -nc -1*
* I wonder if anyone could help me fix this charge issue. *
*Thank you *
*Arooma*
_______________________________________________
AMBER mailing list
AMBER.ambermd.org
http://lists.ambermd.org/mailman/listinfo/amber
- chemical/x-pdb attachment: par.pdb
- application/octet-stream attachment: lig.am1bcc.mol2
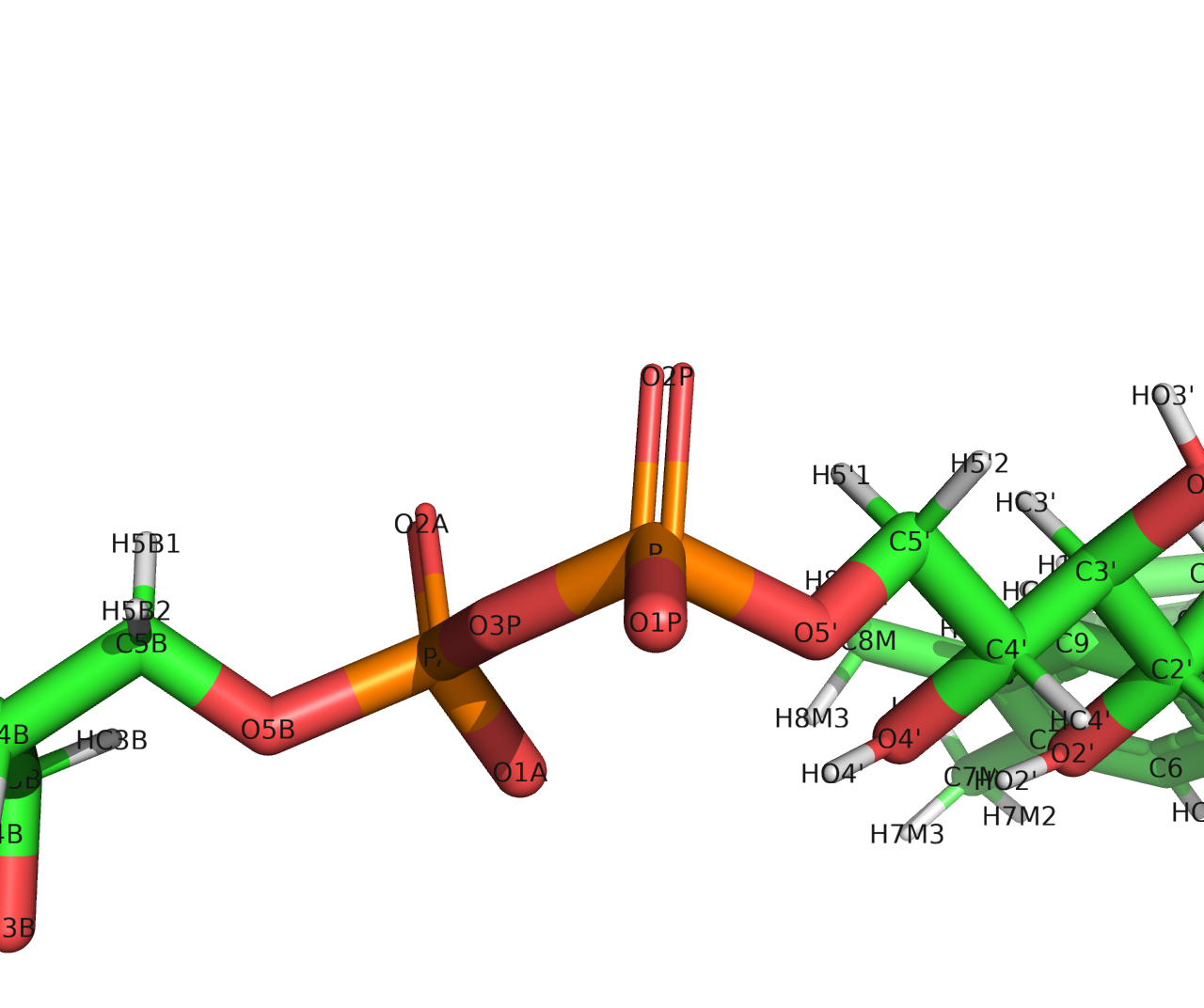
(image/png attachment: P04.png)
