From: Surabhi Singh 24933024 via AMBER <amber.ambermd.org>
Date: Wed, 6 May 2026 12:07:42 +0530
Hello Amber Team,
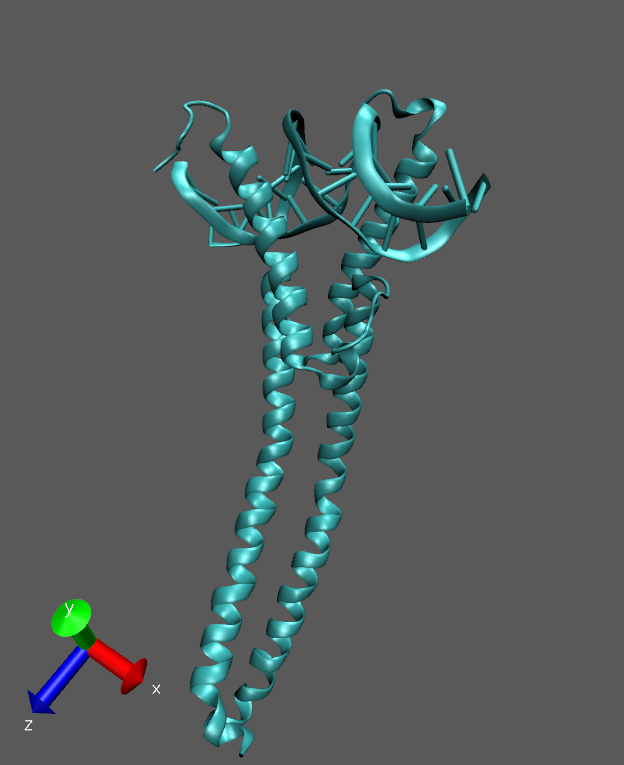
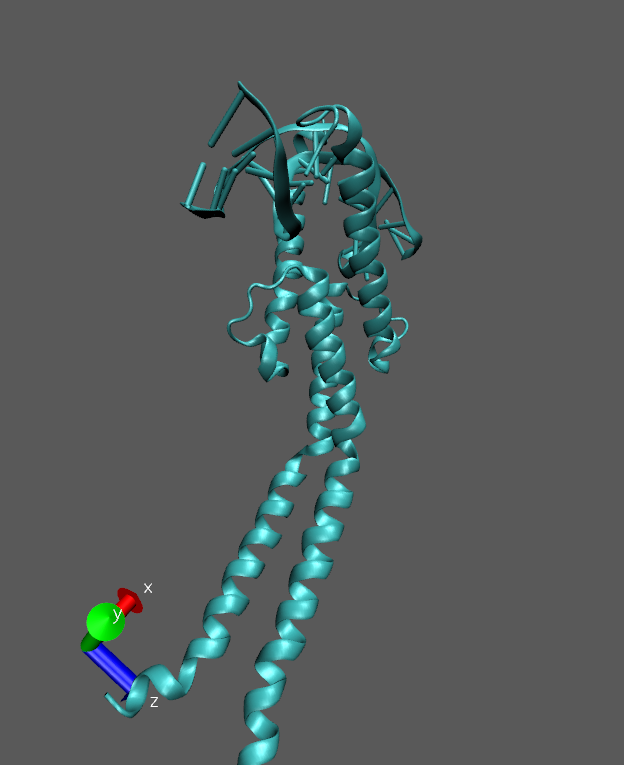
I am working on a protein–DNA complex (bHLH transcription factor). During
the production run, I observe that the longer helical region becomes highly
flexible and shows noticeable bending near the center.
Protocol followed:
1. Forcefield used : Amber: ff19SB, OL24, TIP3P water model
2. Salt conc: 150mM Nacl
3. Energy minimization
4. Three equilibration stages (~1.15 ns total, first two with restrain
and last one without restrain)
5. Production MD: 250 ns (trajectory appears stable overall)
The total and potential energies remain stable with no significant
deviations, yet the helix undergoes bending.
Question:
1. How should this bending be interpreted despite stable energy profiles?
Is this expected behavior (e.g., intrinsic flexibility), or could it
indicate an issue with the setup?
Thank you for your guidance.
Thanks and Regards
Surabhi Singh
Ph.D. Computational Biology
Biosciences and Bioengineering, IIT Roorkee
_______________________________________________
AMBER mailing list
AMBER.ambermd.org
http://lists.ambermd.org/mailman/listinfo/amber
Received on Wed May 06 2026 - 00:00:02 PDT