Date: Thu, 10 Jun 2021 11:07:38 +0200
Dear Prof. David Case,
Thanks for the suggestion about the alignframe and setframe.
I tried this approach, without the common base "C" to connect two RNA
duplexes. but the alignment issue is not solved. Please see attached NAB
script and image.
I tried alignframe+setframe on individual nucleotides, it works on them.
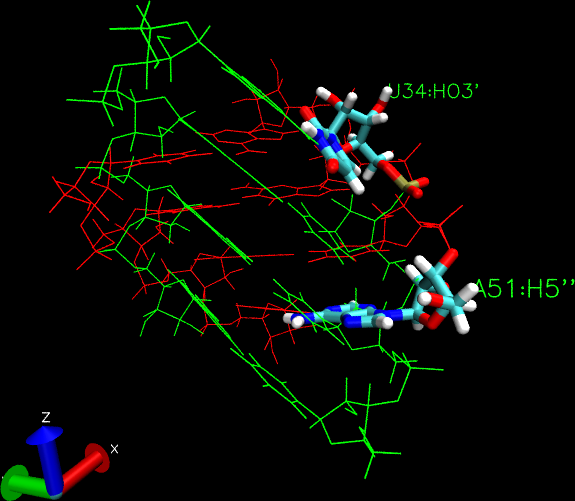
But, for RNA duplexes, I am facing issues. Two RNA duplexes get entangled
with this approach (please see attached image, bases used for
alignframe+setframe are shown in licorice).
Do I need to try different atom expressions to define origin and X, Y axes
to make it work? Please let me know your thoughts.
Best Regards,
Mandar Kulkarni
On Tue, Jun 8, 2021 at 7:07 PM David A Case <david.case.rutgers.edu> wrote:
> On Mon, Jun 07, 2021, Mandar Kulkarni wrote:
> >
> >superimpose( m_align,"1,10" , m_ref, "5,6");
>
> The second and fourth arguments to superimpose should be atom expressions,
> not just strings. Atom expressions have three subsections, separated by
> colons. The superimpose command requires that the two atom expressions
> return the same number of atoms, and that the two atom lists should be
> matched in an ordered fashion. If the two molecules are not chemically
> identical, one has to be very careful in choosing atom expressions that
> have
> the correct number of atoms in the correct order.
>
> I think what you are looking for is a setframe/alignframe pair: that will
> align two molecules together in a way that can work even when they have
> different chemistries.
>
> ...hope this helps...dac
>
>
> _______________________________________________
> AMBER mailing list
> AMBER.ambermd.org
> http://lists.ambermd.org/mailman/listinfo/amber
>
_______________________________________________
AMBER mailing list
AMBER.ambermd.org
http://lists.ambermd.org/mailman/listinfo/amber
(image/png attachment: align.png)
- application/octet-stream attachment: alignframe.nab
