Date: Wed, 17 Oct 2018 17:35:22 +0530
On Wed, Oct 17, 2018 at 5:25 PM Chetna Tyagi <cheta231.gmail.com> wrote:
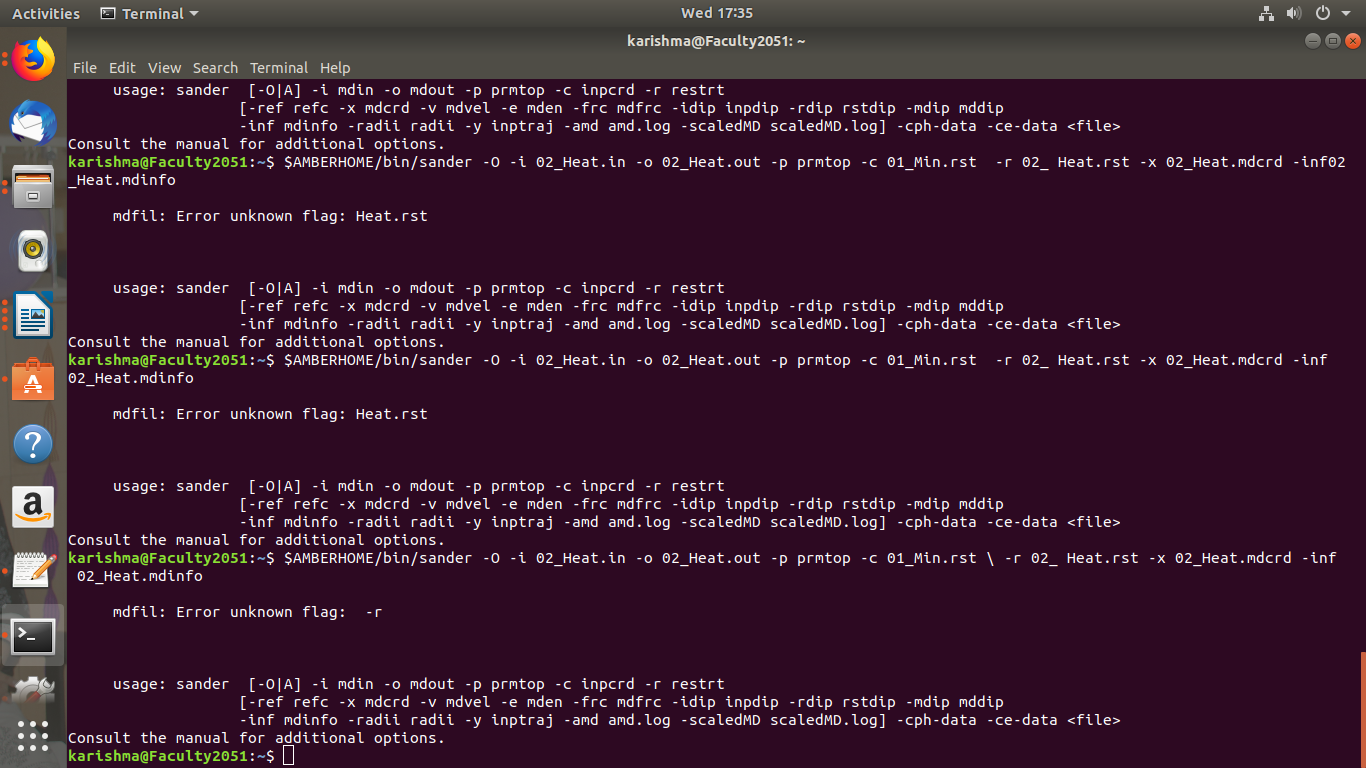
> Please try once without the slash '\' sign...
>
> And it should be -inf Heat.mdinfo...you have written -inf0
>
> On Wed, Oct 17, 2018 at 1:51 PM Charu Sharma (JRF) <
> charu.sharma.lnmiit.ac.in> wrote:
>
> > NOW THIS ERROR I AM GETTING
> >
> > On Wed, Oct 17, 2018 at 5:07 PM Chetna Tyagi <cheta231.gmail.com> wrote:
> >
> > > Hi Karishma,
> > >
> > > The screenshot you have attached shows the error "unknown flag -0". It
> is
> > > -O the alphabet.
> > >
> > > You seem to have typed zero, i.e. '-0' instead of '-O'. If you have
> > already
> > > corrected this, then ignore this email.
> > >
> > > -Chetna
> > >
> > > On Wed, Oct 17, 2018 at 1:30 PM Charu Sharma (JRF) <
> > > charu.sharma.lnmiit.ac.in> wrote:
> > >
> > > > Respected sir,
> > > > i have a doubt in this command can you please clarify this? what
> > > actuality
> > > > is the mistake i am doing
> > > >
> > > > On Sun, Oct 14, 2018 at 1:33 PM Elvis Martis <
> elvis.martis.bcp.edu.in>
> > > > wrote:
> > > >
> > > > > Hi,
> > > > > The command you used to run the SANDER job, you have used "-c" flag
> > > > twice,
> > > > > hence, the error of unknown flag.
> > > > > And second, please ensure that "01_min.in" file is in the working
> > > > > directory. The error indicates it's missing or not readable.
> > > > > Best Regards
> > > > > Elvis Martis
> > > > > Mumbai, INDIA.
> > > > >
> > > > > ________________________________________
> > > > > From: Charu Sharma (JRF) <charu.sharma.lnmiit.ac.in>
> > > > > Sent: 13 October 2018 16:16
> > > > > To: amber.ambermd.org
> > > > > Subject: Re: [AMBER] Regarding Leap
> > > > >
> > > > > Hello sir,
> > > > > I am sending you the screen shot this [problem is showing by
> running
> > > all
> > > > > commands
> > > > >
> > > > > On Sat, Oct 13, 2018 at 1:04 PM Elvis Martis <
> > elvis.martis.bcp.edu.in>
> > > > > wrote:
> > > > >
> > > > > > HI
> > > > > > Can you send the exact command you used, the input file and the
> > error
> > > > > > message?
> > > > > >
> > > > > > Best Regards
> > > > > > Elvis Martis
> > > > > > Mumbai, INDIA.
> > > > > >
> > > > > > ________________________________________
> > > > > > From: Charu Sharma (JRF) <charu.sharma.lnmiit.ac.in>
> > > > > > Sent: 13 October 2018 11:28
> > > > > > To: amber.ambermd.org
> > > > > > Subject: Re: [AMBER] Regarding Leap
> > > > > >
> > > > > > Hello sir,
> > > > > > I would like to know ,i have created the AMBER MD sander input
> > > files .
> > > > > > While following the step of RUN AMBER MD sander ,run the
> > minimization
> > > > of
> > > > > > alanine dipeptide with sander but it showing error of unknown
> flag
> > > why
> > > > ?
> > > > > > when i gave enter command for it .
> > > > > >
> > > > > >
> > > > > >
> > > > > > On Wed, Oct 10, 2018 at 8:48 PM Elvis Martis <
> > > elvis.martis.bcp.edu.in>
> > > > > > wrote:
> > > > > >
> > > > > > > HI,
> > > > > > > Once you have installed AMBER.
> > > > > > > type the following in your terminal
> > > > > > > source $AMBERHOME/amber.sh
> > > > > > > or
> > > > > > > source $AMBERHOME/amber.csh
> > > > > > > and then type
> > > > > > > xleap
> > > > > > >
> > > > > > > Best Regards
> > > > > > > Elvis Martis
> > > > > > > Mumbai, INDIA.
> > > > > > >
> > > > > > > ________________________________________
> > > > > > > From: Charu Sharma (JRF) <charu.sharma.lnmiit.ac.in>
> > > > > > > Sent: 10 October 2018 16:31
> > > > > > > To: amber.ambermd.org
> > > > > > > Subject: Re: [AMBER] Regarding Leap
> > > > > > >
> > > > > > > Hello sir,
> > > > > > > I want to know how to run the command of xleap in ubuntu while
> > > > working
> > > > > in
> > > > > > > linux.
> > > > > > >
> > > > > > > On Tue, Oct 9, 2018 at 11:13 AM Elvis Martis <
> > > > elvis.martis.bcp.edu.in>
> > > > > > > wrote:
> > > > > > >
> > > > > > > > Hi Charu/Karishma,
> > > > > > > > It is highly recommended that you read the AMBER manual (
> > > > > > > > http://ambermd.org/doc12/Amber18.pdf), and also, visit the
> > > amberMD
> > > > > web
> > > > > > > > page (http://ambermd.org/FileFormats.php) that has all your
> > > > > answers.
> > > > > > > > Here, you will find information on FF14SB (
> > > > > > > > http://pubs.acs.org/doi/abs/10.1021/acs.jctc.5b00255)
> > > > > > > > Leaprc.ff14SB is the file to source Amber14 force field
> > > parameters.
> > > > > > > > FYI, from AMBER16 onwards this is "leaprc.protein.ff14SB"
> > > > > > > >
> > > > > > > > Hope this helps
> > > > > > > >
> > > > > > > > Best Regards
> > > > > > > > Elvis Martis
> > > > > > > > Mumbai, INDIA.
> > > > > > > >
> > > > > > > > ________________________________________
> > > > > > > > From: Charu Sharma (JRF) <charu.sharma.lnmiit.ac.in>
> > > > > > > > Sent: 09 October 2018 10:03
> > > > > > > > To: amber-subscribe.ambermd.org; Dr. Ashok Garai;
> > > > AMBER.ambermd.org
> > > > > > > > Subject: [AMBER] Regarding Leap
> > > > > > > >
> > > > > > > > Hello everyone,
> > > > > > > > This is karishma , i want to know basic difference between
> > AMBER
> > > > > > topology
> > > > > > > > file extension, coordinates file extension as well as about
> > > > > > > Leaparc.ff14SB.
> > > > > > > >
> > > > > > > > Thanking you,
> > > > > > > >
> > > > > > > >
> > > > > > > > Regards
> > > > > > > >
> > > > > > > > karishma Sharma
> > > > > > > > _______________________________________________
> > > > > > > > AMBER mailing list
> > > > > > > > AMBER.ambermd.org
> > > > > > > > http://lists.ambermd.org/mailman/listinfo/amber
> > > > > > > >
> > > > > > > > _______________________________________________
> > > > > > > > AMBER mailing list
> > > > > > > > AMBER.ambermd.org
> > > > > > > > http://lists.ambermd.org/mailman/listinfo/amber
> > > > > > > >
> > > > > > > _______________________________________________
> > > > > > > AMBER mailing list
> > > > > > > AMBER.ambermd.org
> > > > > > > http://lists.ambermd.org/mailman/listinfo/amber
> > > > > > >
> > > > > > > _______________________________________________
> > > > > > > AMBER mailing list
> > > > > > > AMBER.ambermd.org
> > > > > > > http://lists.ambermd.org/mailman/listinfo/amber
> > > > > > >
> > > > > > _______________________________________________
> > > > > > AMBER mailing list
> > > > > > AMBER.ambermd.org
> > > > > > http://lists.ambermd.org/mailman/listinfo/amber
> > > > > >
> > > > > > _______________________________________________
> > > > > > AMBER mailing list
> > > > > > AMBER.ambermd.org
> > > > > > http://lists.ambermd.org/mailman/listinfo/amber
> > > > > >
> > > > >
> > > > > _______________________________________________
> > > > > AMBER mailing list
> > > > > AMBER.ambermd.org
> > > > > http://lists.ambermd.org/mailman/listinfo/amber
> > > > >
> > > > _______________________________________________
> > > > AMBER mailing list
> > > > AMBER.ambermd.org
> > > > http://lists.ambermd.org/mailman/listinfo/amber
> > > >
> > >
> > >
> > > --
> > > Best wishes
> > > Chetna
> > > _______________________________________________
> > > AMBER mailing list
> > > AMBER.ambermd.org
> > > http://lists.ambermd.org/mailman/listinfo/amber
> > >
> > _______________________________________________
> > AMBER mailing list
> > AMBER.ambermd.org
> > http://lists.ambermd.org/mailman/listinfo/amber
> >
>
>
> --
> Best wishes
> Chetna
> _______________________________________________
> AMBER mailing list
> AMBER.ambermd.org
> http://lists.ambermd.org/mailman/listinfo/amber
>
_______________________________________________
AMBER mailing list
AMBER.ambermd.org
http://lists.ambermd.org/mailman/listinfo/amber
(image/png attachment: Screenshot_from_2018-10-17_17-35-08.png)
