Date: Wed, 17 Oct 2012 21:27:22 -0400
Next time, it will be simpler if you attach the pdb file rather than
copying it into the text.
I built your structure (pdb attached), but it needs some modification.
I used our current/old site (glycam.org). Perhaps the structure will
be useful, even in its current state.
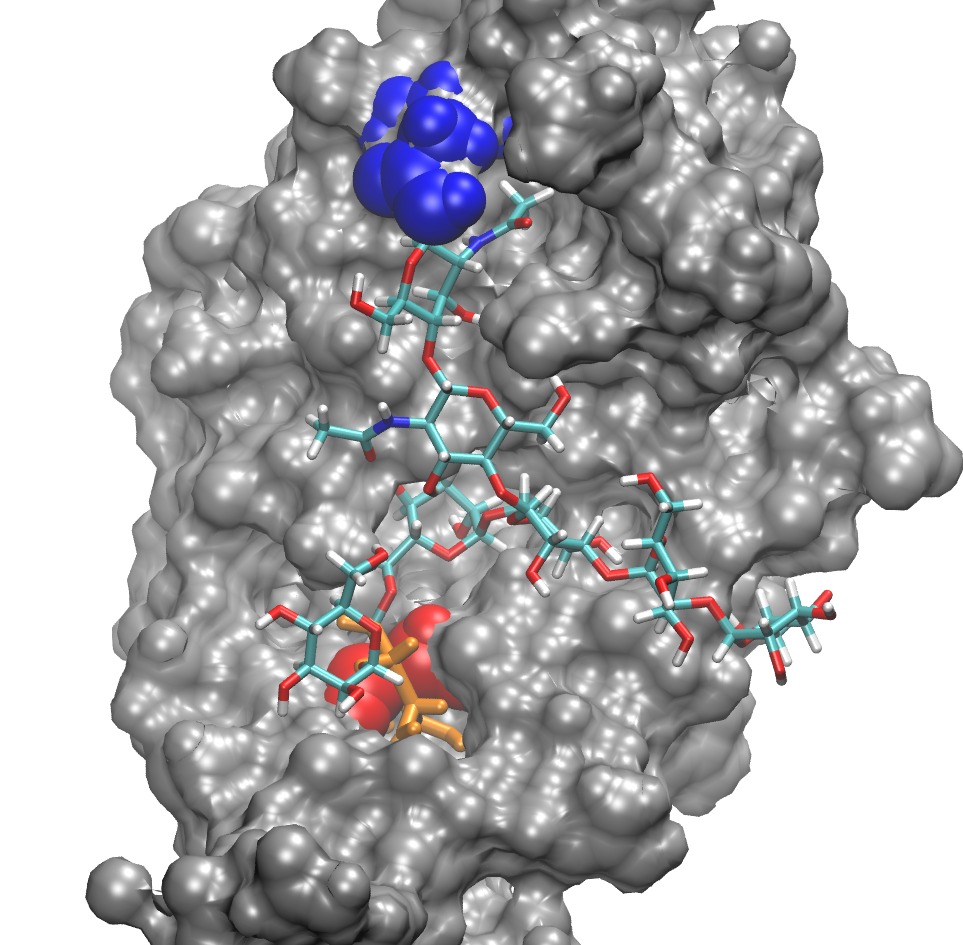
See the attached image. The glycan (stick representation) has wrapped
itself nicely around the protein (gray surface), and seems to fit well
into the grooves. I don't know, however, if the "grooves" would
happen naturally or are an unrealistic result of minimization forces.
The ASN where it attaches is in blue. The biggest problem is with one
residue (orange) that was in a bad position to start with. As a
result, in this minimized structure, the C-CA bond in one of the amino
acids (red) spanned the ring (orange). That is, the C-CA bond sticks
through the ring. We are working on automated algorithms for working
things like this out, but I believe you could alter the
DManpa1-6DManpa omega angle to relieve the strain.
One thing our new site will do is to allow you to alter glycosidic
angles. But, while trying to build the structure at that site
(dev.glycam.org), I found some bugs. We'll get those fixed as soon as
we can.
Please double-check my glycan. This is what I added:
DManpa1-2DManpa1-6[DManpa1-3]DManpa1-6[DManpa1-2DManpa1-3]DManpb1-4DGlcpNAcb1-4DGlcpNAcb1-OME
By the way, the structural issues explain why the minimization took
longer than usual. This structure took a little over 200 seconds,
about twice the predicted time. We see this sort of thing pretty
often. Like I said, we are working hard to come up with good
automated ways to deal with challenging structures such as these.
-- :-) Lachele Lachele Foley CCRC/UGA Athens, GA USA
_______________________________________________
AMBER mailing list
AMBER.ambermd.org
http://lists.ambermd.org/mailman/listinfo/amber
- chemical/x-pdb attachment: blftst_10_17_2012_212_MIN_formatted.pdb
(image/jpeg attachment: glycoprotein_ring_strain_dat.jpg)
